FAQs
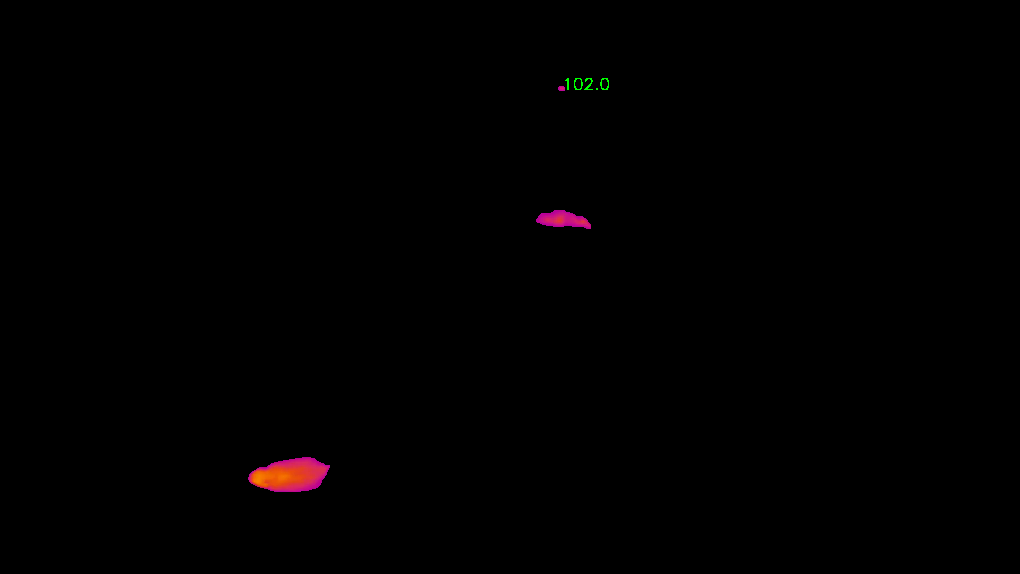
Threshold based object segmentation
Added track bars for users to select HSV values to segment ROIs in the provided video.
python annolid/main.py -v /path/to/my_video.mp4 --segmentation=threshold

Convert WMV format to mp4 format using ffmpeg
ffmpeg -i /path/to/my_video.wmv -c:a aac /path/to/my_video.mp4
Save the extracted frames to a user selected output directory
If not selected, it will save the extracted frames to a folder named with the video name without extension. For example, if the input video path is /path/to/my_video.mp4, the extracted frames will be saved in the folder /path/to/my_video. The output directory is provided, the extracted frames will be saved /path/to/dest/my_video.
cd annolid
python main.py -v /path/to/my_video.mp4 --extract_frames=20 --to /path/to/dest --algo=uniform
How to track multiple objects in the video?
YOLO-based tracking has been removed from Annolid.
How to convert labelme labeled dataset to COCO format?
Use the GUI export dialog:
Open Annolid.
Go to Convert -> LabelMe -> COCO.
Select your LabelMe annotation directory.
Optionally set output directory and labels file.
Choose train split and output mode (Segmentation or Keypoints), then click OK.
You can also run the CLI:
python annolid/main.py \
--labelme2coco=/path/to/my_labeled_images \
--to /path/to/my_dataset_coco \
--labels=/path/to/my_labels.txt
The dataset is structured as:
../../datasets/mydataset_coco/
├── data.yaml
├── annotations_train.json
├── annotations_valid.json
├── train
│ ├── annotations.json
│ └── JPEGImages
│ ├── 00000444.jpg
└── valid
├── annotations.json
└── JPEGImages
├── 00000443.jpg
Convert the tracking results csv file to Glitter2 csv format
The result csv file named as tracking_results_nix.csv in the folder as provided in –to option.
python annolid/main.py -v /path/to/my_video.mkv --tracks2glitter /path/to/tracking_results.csv --to /path/to/results_dir/
Convert the keypoint annotations to labelme format
python annolid/main.py --keypoints2labelme /path/to/mouse_m7s3/ --keypoints /path/to/mouse_m7s3/CollectedData_xxxx.h5